This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is transcriptomics?
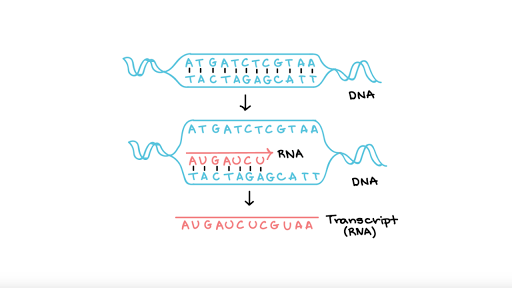 Transcription
Transcription
Transcriptomics is the study of all the RNA transcripts present in the cell. [1] Since RNA is transcribed from DNA, analyzing the amount of RNA transcripts present in a specific tissue can give us an understanding for which genes are being expressed and to what degree. [1] Figuring out what other genes are upregulated or downregulated along with a gene of interest can help us determine what other genes are affected by that gene of interest. [1] The main methods used to explore the transcriptome are microarray analysis and RNA-sequencing (RNA-seq).
What is RNA-sequencing?
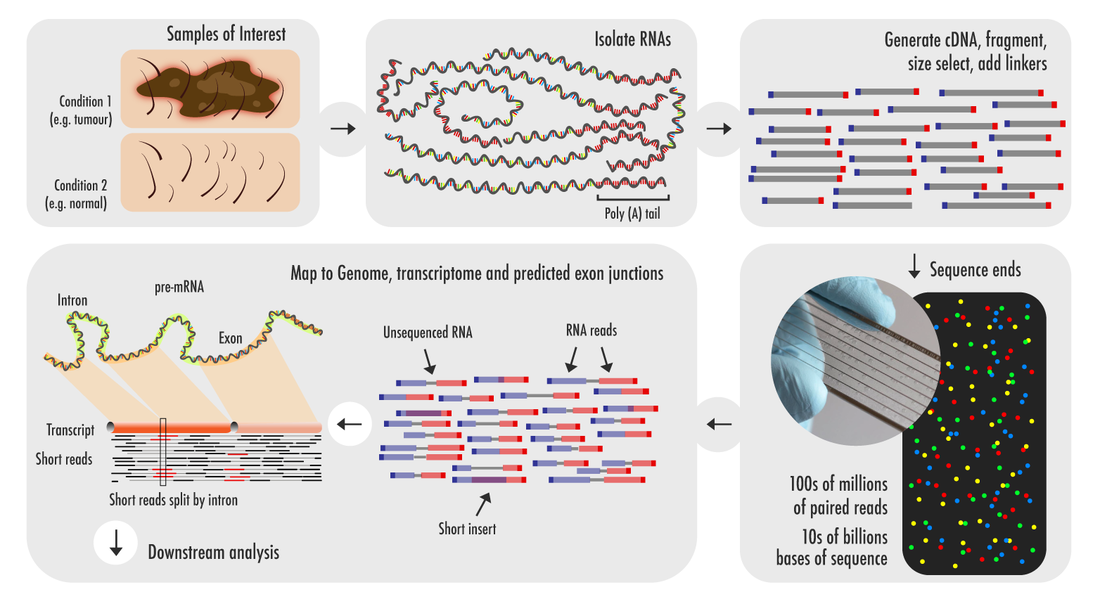
RNA-seq can determine the quantity and sequence of RNA transcripts present in the cell using next generation sequencing (NGS). [2] With this technique we can discover which genes are turned on in a cell along with their expression level and when they are active in response to other variables or disease states. [2] The process begins with isolating the RNAs from a sample of interest and generating cDNA via RT-PCR. These cDNAs are then fragmented and adapters are added to the ends that permit sequencing. A figure describing and overview of RNA-seq is provided below.
Conclusion
Transcriptomics is a vital method for exploring complex diseases such as PCOS. Using The Human Protein Atlas I found that StAR is expressed mainly in the adrenal glands, ovaries, and testes. RNA-seq was a useful tool in another study that looked at the expression of StAR and related genes in prenatally androgenized rats expressing PCOS-like symptoms. [3] Higher mRNA expression of StAR has been linked to PCOS as a contributing factor to increased testosterone levels in the ovarian cells. [3,4] Further exploration of what other genes are upregulated and downregulation in a PCOS model or patients will elucidate how StAR expression becomes increased.
References
[1] Transcriptome. (2015, August 27). Retrieved March 14, 2019 from https://www.genome.gov/13014330/
[2] RNA-seq: Basics, Applications and Protocol. (2018, April 6). Retrieved March 14, 2019 from https://www.technologynetworks.com/genomics/articles/rna-seq-basics-applications-and-protocol-299461
[3] Jahromi, M. S. et al. (2016). Elevated expression of steroidogenesis pathway genes; CYP17, GATA6 and StAR in prenatally androgenized rats. Gene. 593(1):167-171.
[4] Kahsar-Miller, M. D. et al. (2001). Steroidogenic acute regulatory protein (StAR) in the ovaries of healthy women and those with polycystic ovary syndrome. American Journal of Obstetrics and Gynecology. 185(6):1381-1387.
[1] Transcriptome. (2015, August 27). Retrieved March 14, 2019 from https://www.genome.gov/13014330/
[2] RNA-seq: Basics, Applications and Protocol. (2018, April 6). Retrieved March 14, 2019 from https://www.technologynetworks.com/genomics/articles/rna-seq-basics-applications-and-protocol-299461
[3] Jahromi, M. S. et al. (2016). Elevated expression of steroidogenesis pathway genes; CYP17, GATA6 and StAR in prenatally androgenized rats. Gene. 593(1):167-171.
[4] Kahsar-Miller, M. D. et al. (2001). Steroidogenic acute regulatory protein (StAR) in the ovaries of healthy women and those with polycystic ovary syndrome. American Journal of Obstetrics and Gynecology. 185(6):1381-1387.
Header image: http://bigbangpage.com/wp-content/uploads/2017/03/?SD